Risks and Limitations of AI in the Life Sciences
After nearly 20 years focused on mathematics, machine learning, and AI ethics, I went back to school and completed a Masters in Microbiology-Immunology. Last month, Kamayani Gupta, co-founder of KAMI Think Tank, hosted me for a Q&A about risks and limitations of AI in the life sciences. What follows below is an edited and shortened version of our conversation. Or watch our full-length discussion in the video here:
Kamayani: My first question: we’re seeing AI applied quite a bit throughout life sciences, and there’s a lot of hype versus what’s actually being built properly. Where do you think confidence is running ahead of actual scientific understanding?
Rachel: This is a big issue. I’m excited about AI — I work for an AI startup — but the confidence and hype often outpace reality. One big concern is the assumption that we already have all the data we need, and just need to throw it into a model for amazing outputs. What worries me is that we may be underinvesting in thinking about new types of data. The type and quality of data really sets limits on the quality of results. With medicine, I see this assumption that patient records and electronic health records will unlock breakthroughs — whereas in many cases we need new assays, new biomarkers we haven’t discovered yet. There’s still a huge need for bench and lab research, and I’m worried that is not getting the funding that AI applications are.
AlphaFold is probably the biggest success story, and it is genuinely impressive — but people lose sight of the fact that the Protein Data Bank (PDB) and the Critical Assessment of Structure Prediction (CASP) competition are what made it possible. The PDB started in the 1970s on magnetic tape sent through the mail. CASP was thoughtfully structured and has been running since the 1990s. The AlphaFold team’s innovations are truly impressive, but they needed the right type of high-quality data that was a good fit for the problem. In many cases the data isn’t the right fit, and people just say, “this is what we have, let’s go for it.”
Kamayani: That’s such an important example — CASP almost lost its funding last year, and it took people calling out how critical that program and the PDB were to AlphaFold’s existence. It’s decades of work, not a company spun up two years ago. The other thing that always strikes me is how hard it is to evaluate these models without deep biological expertise. Metrics can look really strong from the outside without the biology actually making sense. So when AI systems in biology are wrong, who usually discovers that, and where does ownership lie for these new systems being built today?
Rachel: That’s exactly the right question. We connected after you read my blog post about the enzyme classification paper, which is a really important case study. Published in Nature Communications, the team used 22 million enzymes to predict enzyme function from amino acid sequences. On its own, the paper seemed sound — they had training, validation, and test sets, and afterwards applied their model to 450 enzymes with unknown functions, checking three in the lab.
What happened is a microbiologist, Dr. Valérie de Crécy-Lagard, who had studied one of those enzymes for over a decade, recognized that the paper’s conclusion about it was simply wrong — she had already disproven it in the lab. When she dug into the other results, she found hundreds of errors. 135 of the “novel” enzymes already appeared in UniProt — significant data leakage. Some results were blatantly implausible, like attributing mycothial synthase to an enzyme in E. coli, which doesn’t synthesize mycothial. And 12 different enzymes were assigned the same narrow function, pointing to overfitting.
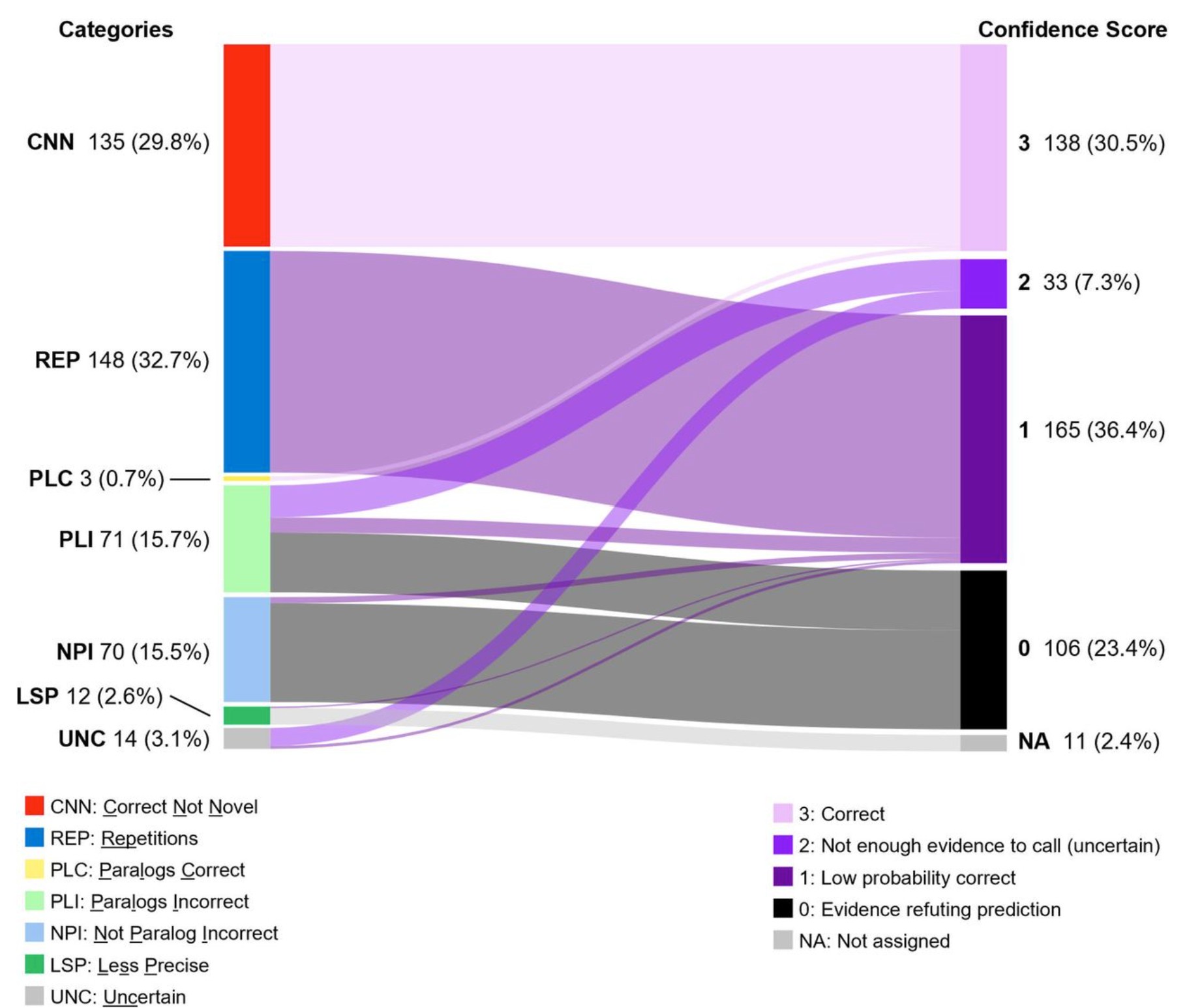
None of this would have been caught without someone with her specific expertise happening to read the paper. She then had enormous difficulty getting her rebuttal published — she contacted the authors, contacted Nature Communications, assembled a team, and went through multiple rejections. That really illustrates the incentive problem: the exciting AI result gets into the prestigious journal, and refuting it is a much harder road.
Kamayani: That’s striking — both the errors and how hard it was to correct the record. It raises the question of ownership: when something goes wrong, does responsibility lie with the company that built the model, the company that used it, or the governing agency that assessed it?
Rachel: It occurs at so many levels. This case points to the need for deep integration with domain experts — microbiologists closely involved throughout. It also points to a field that simply doesn’t reward error-checking work, so it falls through the cracks. The rebuttal paper was fascinating and important research, but there’s no funding, support, or recognition for that kind of work.
And it can be genuinely hard to construct a training/validation/test split that avoids data leakage. We saw with the CASP competition that it took a dedicated committee with real funding to do it well. Individual teams are often under-resourced, and these methodological questions just don’t get the attention that model architecture does.
Kamayani: And I think you’ve already answered what I was going to ask next — what incentives in AI research or deployment worry you most?
Rachel: There is a paper I love called “Everyone Wants to Do the Model Work, Nobody Wants to Do the Data Work”– a great title that most of us in data science can relate to. The researchers interviewed over 60 machine learning practitioners across three continents and talked about data cascades: what can go wrong in high-stakes ML applications.
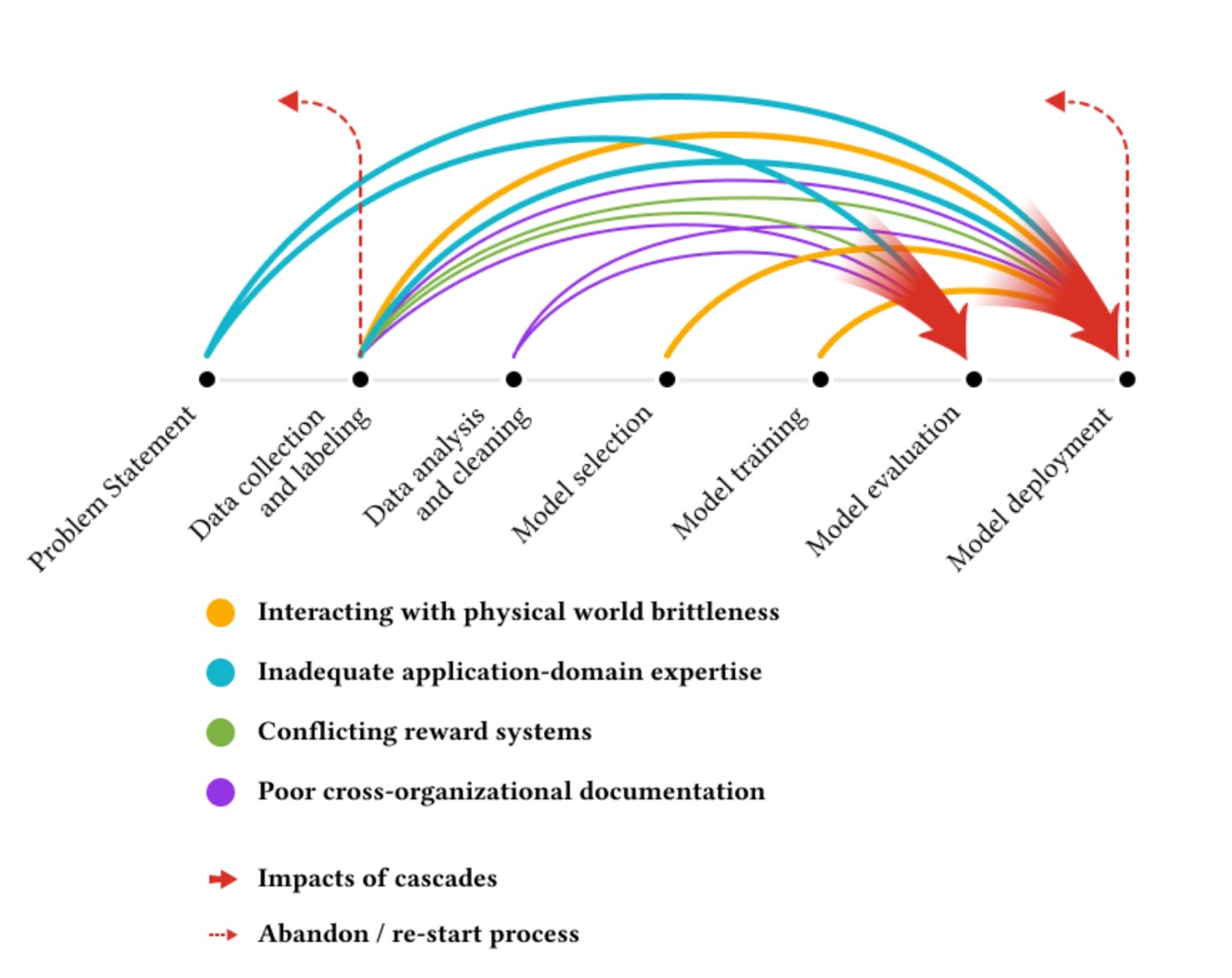
In so many cases, people in the field were asked to collect extra data but weren’t given extra pay or time to do so. Measurement systems would change in the field and that wouldn’t make it back to the computer lab where people were building models. There’s a case where an anti-poaching model, when they got to deployment, was producing results the anti-poaching teams said were incorrect. It turned out there were issues with the underlying dataset. If there had been more integration across roles earlier, that could have been prevented. A lot of it comes down to ensuring collaboration throughout the process.
Kamayani: Large datasets and big models lend this air of authority — bigger is better seems ingrained in us. But when does scale become misleading rather than reassuring?
Rachel: Scale can often be misleading — especially when data has systematic biases rather than random noise, when particular types of data are missing, or when the underlying paradigm is incorrect. An example: early in COVID, there was an app in the UK called Zoe, originally a diet and nutrition tracker that was quickly modified to track COVID cases. It was designed around a short-term respiratory virus, so when people developed Long COVID, they couldn’t log their symptoms properly. Neurological symptoms, fatigue, brain fog — none of those were included in the preset options. People were hand-typing symptoms and having to re-enter them every day for months, because the app wasn’t built for long-term tracking.
This data was then used in research studies on long COVID prevalence, with faulty assumptions like “people stopped using the app, so they must have recovered.” I credit researcher Hannah Davis for surfacing this issue– the data simply wasn’t designed for that purpose. Scaling it to even more users wouldn’t have helped. It needed a fundamentally different design to answer those questions.
Kamayani: And long COVID is so diverse — each person is affected differently — so even with a massive dataset, if the collection mechanism is wrong, you end up with something chaotic and noisy the moment you try to build a model on top of it.
Rachel: Particularly when you lose sight of the fact that no matter what data you’re gathering, there are decisions that go into the design of how you gather it: what you include, what questions you ask — and those shape what the data set looks like. People often think data is objective truth, but it’s constructed through a series of decisions that really matter.
Another important point from that case study: there were patients reaching out to the Zoe app creators saying this isn’t meeting my needs, and that feedback was not incorporated. That really highlights the importance of listening to patients, because they have a firsthand perspective on how a tool is failing them.
Kamayani: A lot of times that feedback loop doesn’t even get generated. And as more people use AI modeling, incorrect predictions that get published or fed into databases don’t just sit there — they become training data for the next model.
Rachel: I worry this happens with diseases that are underdiagnosed or have diagnostic delays — the model sees it as rarer than it is and therefore less likely. Take lupus, where the average time to diagnosis is six to eight years. Consider how many patients have not received an accurate diagnosis yet or who give up before ever finding one. This leads to incomplete and missing medical data. That’s the data getting fed into these models, and you get self-reinforcing feedback loops.
Kamayani: So if teams genuinely want to reduce harm to patients, what fundamental practices have to change — even if that means moving more slowly, which I know is counterintuitive to the “AI moves faster” messaging?
Rachel: Go slow to go far. I think it’s really important that we continue investing in research focused on underlying causal mechanisms. Our current AI systems are doing a fuzzy interpolation between existing data points — valuable, but because of that, they won’t give us something truly outside the scope of the training data. We still need research where new paradigms or different causal mechanisms are required.
I’ll cite Arijit Chakravarty, who has worked across pharmaceutical development and coined the concept of “frankencells.” When people pull together pathways from different papers — something AI and mathematical modeling encourages — you can end up with diagrams that would never all occur in a single cell. In cancer research, there are published pathways where each individual arrow is correct, but they wouldn’t all happen in the same cell. That’s the temptation with AI: throwing results together without thinking about the underlying mechanism. He argues cancer development should be understood as an evolutionary process with randomness, not a circuit diagram.
Beyond that, continuing to invest in bench science matters. And then much of what we’ve discussed comes down to meaningful, ongoing collaboration: domain experts at every stage of data collection and processing, model development, patients, and clinicians who will actually use the tool.
Kamayani: One last question before we go to the audience: what’s been a really interesting or innovative use case you’ve seen in life sciences recently?
Rachel: T-cell binding is something I’ve done a deep dive on. It’s a field where there’s still a lot of work to be done — there’s even an ongoing competition around it, which I find fascinating. The way well-structured competitions can push innovation still excites me. We’ve seen it with AlphaFold and AlexNet, both arising out of competitions that had been running for years.
These competitions also force people to be explicit about their data, and that’s my big caveat with AI. You need to be clear: this is the data I’m using, these are the constraints, these are the biases, this is what wasn’t collected. I love Timnit Gebru’s Data Sheets for Datasets paper: being specific about what data was collected, what the appropriate uses are, and where it wouldn’t apply. When you use machine learning within clear parameters, it’s quite valuable.
Kamayani: A lot of people we work with are trying to upskill, often biologists moving into AI. What technique or tactic would you recommend for learning these hard topics?
Rachel: I co-founded fast.ai, which still has valuable free courses on AI and deep learning. Now with Answer AI we run paid courses around a style of problem solving where you use AI to break things down into small pieces you can understand, keeping yourself in the loop to really understand the problem.
Kamayani: An audience question: “Are there any interesting AI tools we should know about?”
Rachel: My biased answer: SolveIt, which I’m working on. It’s a Jupyter notebook-like where you can run AI prompts directly within the notebook. One feature I love is that you can edit the AI’s output — so instead of getting in a long argument with AI and polluting the context, you can fix it directly. It’s designed to keep you in the loop rather than going off and building huge solutions for you.
Kamayani: Thank you so much, Rachel — every time I speak with you I learn something new. Check out Rachel’s blog and answer.ai. Also, KAMI Think Tank hosts events every month, so join our membership if you’re interested. Thanks everyone, and have a great evening.
Rachel: Thanks so much for hosting! This was a lot of fun.
Related Posts: